Viruses depend on and modulate their hosts’ cellular environments to maximize replication. Studies of viruses can therefore reveal important aspects of host-pathogen interactions and fundamental cell biology. Viruses often modulate host pathways using proteins, but can also express non-coding RNAs whose functions and mechanisms are mostly unknown.
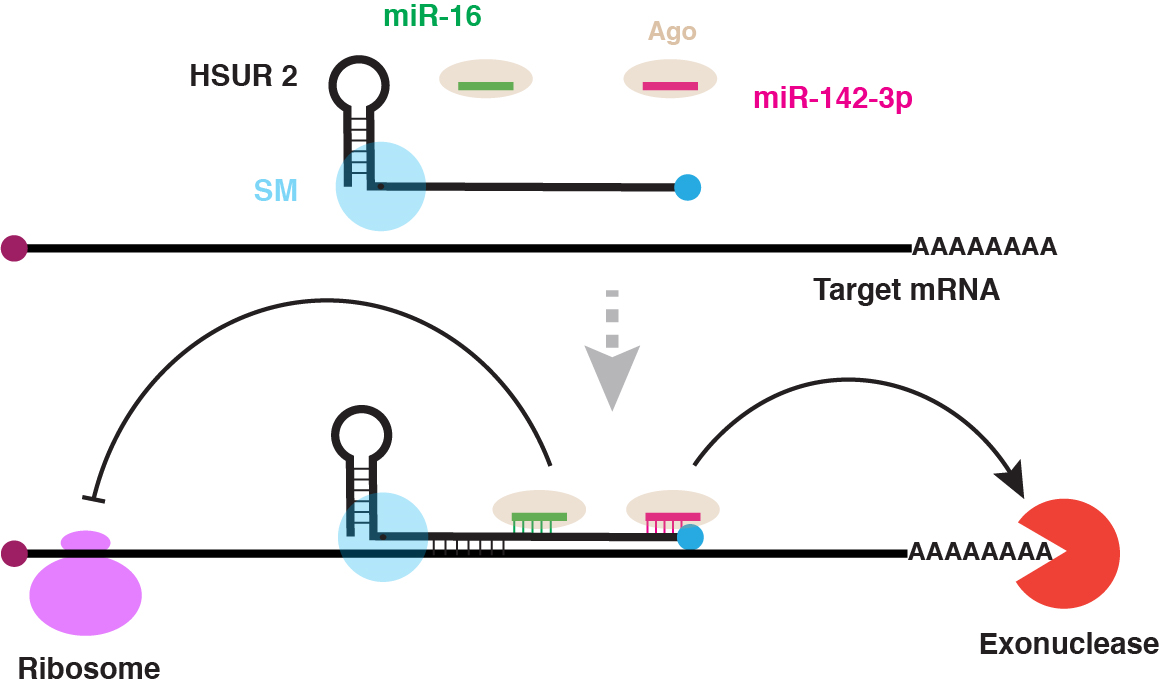
Cazalla and colleagues studied the small RNAs from H. saimiri, a herpesvirus that establishes latency in the T cells of New World primates and can cause aggressive leukemias and lymphomas in non-natural hosts. They showed these RNAs, called HSURs, modulate host gene expression and inhibit host cell death using a novel mechanism in which the HSURs inhibit host mRNAs by tethering them to host miRNAs and the associated degradation and translation inhibition machinery. This mechanism is a completely novel process, not previously observed in cells, and but which promises to lead to a fuller understanding of gene regulation in both infected and uninfected cells.
References:

A viral Sm-class RNA base-pairs with mRNAs and recruits microRNAs to inhibit apoptosis. Gorbea C, Mosbruger T, Cazalla D. Nature. 2017 Oct;550(7675):275.
Press Releases and Media:

University of Utah Health: A Viral “Bait and Switch” Boosts Infection

